
Chromosome conformation capture
Encyclopedia
High-throughput
High-throughput may refer to:* High-throughput computing - a computer science concept * High-throughput screening - a bioinformatics concept* Measuring data throughput - a communications concept...
molecular biology
Molecular biology
Molecular biology is the branch of biology that deals with the molecular basis of biological activity. This field overlaps with other areas of biology and chemistry, particularly genetics and biochemistry...
technique used to analyze the organization of chromosomes in a cell's natural state. Studying the structural properties and spatial organization of chromosomes is important for the understanding and evaluation of the regulation of gene expression, DNA replication
DNA replication
DNA replication is a biological process that occurs in all living organisms and copies their DNA; it is the basis for biological inheritance. The process starts with one double-stranded DNA molecule and produces two identical copies of the molecule...
and repair
DNA repair
DNA repair refers to a collection of processes by which a cell identifies and corrects damage to the DNA molecules that encode its genome. In human cells, both normal metabolic activities and environmental factors such as UV light and radiation can cause DNA damage, resulting in as many as 1...
, and recombination
Genetic recombination
Genetic recombination is a process by which a molecule of nucleic acid is broken and then joined to a different one. Recombination can occur between similar molecules of DNA, as in homologous recombination, or dissimilar molecules, as in non-homologous end joining. Recombination is a common method...
.
One example of chromosomal interactions influencing gene expression is a chromosomal region which can fold in order to bring an enhancer
Enhancer (genetics)
In genetics, an enhancer is a short region of DNA that can be bound with proteins to enhance transcription levels of genes in a gene cluster...
and associated transcription factor
Transcription factor
In molecular biology and genetics, a transcription factor is a protein that binds to specific DNA sequences, thereby controlling the flow of genetic information from DNA to mRNA...
s within close proximity of a gene, as was first shown in the beta-globin locus. Chromosome conformation capture has enabled researchers to study the influences of chromosomal activity on the aforementioned cellular mechanisms. This technology has aided the genetic and epigenetic
Epigenetics
In biology, and specifically genetics, epigenetics is the study of heritable changes in gene expression or cellular phenotype caused by mechanisms other than changes in the underlying DNA sequence – hence the name epi- -genetics...
study of chromosomes both in model organisms and in humans.
Several techniques have been developed from 3C to increase the throughput of quantifying a chromosome’s interactions with other chromosomes and with proteins. All the 3C related technologies are broadly categorized in to four groups. (1) 3C and ChIP version of 3C (ChIP-loop assay), (2) 4C and ChIP version of 4C (enhanced 4C), (3) 5C and 3D assays and (4) Genome conformation capture (GCC) related (Hi-C), ChIP version of GCC as 6C. The application of analyzing DNA segments by microarray
Microarray
A microarray is a multiplex lab-on-a-chip. It is a 2D array on a solid substrate that assays large amounts of biological material using high-throughput screening methods.Types of microarrays include:...
and high-throughput sequencing in the 4C, 5C and Hi-C methodologies has brought the assessment of chromosome interactions to the genome-wide scale.
Chromosome Conformation Capture (3C)
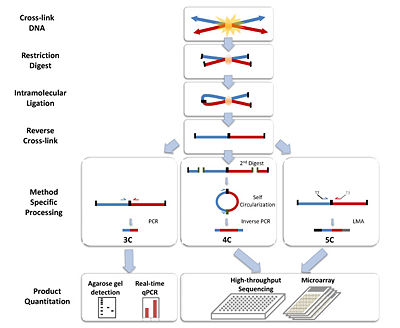
The basic 3C technique has five experimental steps:Step 1 Cross-linking: Addition of formaldehyde
Formaldehyde
Formaldehyde is an organic compound with the formula CH2O. It is the simplest aldehyde, hence its systematic name methanal.Formaldehyde is a colorless gas with a characteristic pungent odor. It is an important precursor to many other chemical compounds, especially for polymers...
results in the cross-linking of DNA segments to proteins and to cross-linking of proteins with each other. This leads to cross-linking of interacting DNA segments (for example cis located promoters to trans located promoters, reveals interactions like the interaction between H enhancer and odorant receptor promoters).
Step 2 Restriction digest
Restriction digest
A restriction digest is a procedure used in molecular biology to prepare DNA for analysis or other processing. It is sometimes termed DNA fragmentation...
: A restriction enzyme
Restriction enzyme
A Restriction Enzyme is an enzyme that cuts double-stranded DNA at specific recognition nucleotide sequences known as restriction sites. Such enzymes, found in bacteria and archaea, are thought to have evolved to provide a defense mechanism against invading viruses...
is added in excess to the cross-linked DNA, separating the non-cross-linked DNA from the cross-linked chromatin. The selection of the restriction enzyme in this step depends on the locus being analyzed and allows the separate analysis of different regulatory elements. Frequently cutting enzymes (4 bp) are used in studying small loci (<10–20 kb), while larger loci demand the use of larger cutters (6 bp). Restriction enzyme selection also has an impact on digestion efficiency of cross-linked DNA.
Step 3 Intramolecular Ligation:
Step 4 Reverse Cross-links: High temperature will result in the reversal of cross-links formed in step 1. The resulting linear DNA fragment has specific restriction ends as well as a central restriction site corresponding to the site of ligation. The pool of these fragments is collectively referred to as the 3C library.
Step 5 Quantitation: Polymerase chain reaction (PCR)
Polymerase chain reaction
The polymerase chain reaction is a scientific technique in molecular biology to amplify a single or a few copies of a piece of DNA across several orders of magnitude, generating thousands to millions of copies of a particular DNA sequence....
uses primers against the site of ligation to semi-quantitatively assess the frequencies of a restriction fragment of interest. Real-time PCR using Taqman
TaqMan
TaqMan probes are hydrolysis probes that are designed to increase the specificity of real-time PCR assays. The method was first reported in 1991 by researchers at Cetus Corporation, and the technology was subsequently developed by Roche Molecular Diagnostics for diagnostic assays and by Applied...
probes (3C-qPCR) provides a more quantitative measurement of the fragment of interest. The Taqman probe and a constant primer hybridize to the restriction fragment that contains the site of contact and one test primer is designed against each neighboring restriction fragments. Together the probe and primers allow for a specific fluorescent signal to be emitted during amplification.
Circularized Chromosome Conformation Capture (4C)
The 4C strategy has a significant advantage over 3C in that only the sequence of one of the site of interest needs to be known. The fragment, known as the “bait”, contains the site that associates with other chromosomal regions. The 4C procedure follows the same steps as in 3C, except additional processing is needed before quantification of the fragments of interest.Steps 1-4: See procedure listed in 3C. 6-bp-cutters are preferred in step 2.
Step 5a Second Restriction Digest: After the reversal of the cross-linked DNA, the restriction fragments are subjected to another round of restriction digest, this time with a frequent cutter that will result in smaller fragments with restriction ends that differ from the central restriction site (ligation junction).
Step 5b Self-circularization: Self-circularization of the DNA fragments is more favored now that they are not bound to other proteins or fragments. Intramolecular ligation occurs to induce the formation of the circular fragments. The pool of circular fragments becomes the 4C library.
Step 5c Inverse PCR
Inverse polymerase chain reaction
Inverse polymerase chain reaction is a variant of the polymerase chain reaction that is used to amplify DNA with only one known sequence...
and Quantitation: Primers are designed against the outer restriction sites of the “bait” sequence, which result in the amplification of the small unknown captured fragment. Large-scale sequencing can be used to sequence the 4C library. Custom microarrays can also be made using probes designed against the adjacent upstream and downstream regions of all genomic sites of the restriction enzyme used in step 2.
Carbon-Copy Chromosome Conformation Capture (5C)
The 5C technique expands from 3C and allows for the parallel analysis of interactions between many selected loci.Steps 1-4: Same as in 3C.
Step 5 Ligation-mediated amplification and Quantitation: Performing multiplex ligation-mediated amplification (LMA)
Multiplex ligation-dependent probe amplification
Multiplex ligation-dependent probe amplification is a variation of the polymerase chain reaction that permits multiple targets to be amplified with only a single primer pair. Each probe consists of a two oligonucleotides which recognise adjacent target sites on the DNA...
after the construction of the 3C library leads requires using multiplex primers that consist of universal primer sequences like T7 and T3 and the ligation junction sequences. They anneal to the 3C fragments and get ligated together with a DNA ligase. Perfect alignment with the 3C template ensures the success of the ligation. The ligated primers serve as templates of which get amplified to generate the 5C library. The use of universal primer sequences mean these 5C fragments can be analyzed on microarrays. The small size (~100 bp) of the 5C fragments is also compatible for analysis using high-throughput sequencing.
ChIP-loop
This method is slightly different from the previous techniques in that the interaction formed between two chromosomal regions is mediated by a bound protein. Like in the 5C methodology, a single DNA site is often considered to interact with multiple other sites. After the cross-linking and digestion, ChIPChromatin immunoprecipitation
Chromatin Immunoprecipitation is a type of immunoprecipitation experimental technique used to investigate the interaction between proteins and DNA in the cell. It aims to determine whether specific proteins are associated with specific genomic regions, such as transcription factors on promoters or...
is performed to pull down the protein bound to the site of interest. Normal 3C procedures are conducted after this step.
Advantages and Disadvantages
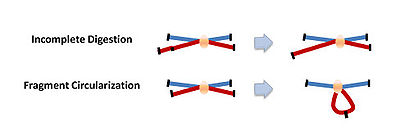
A significant confounding factor in the 3C technology is the frequent random collisions of chromosomal regions to one another, which means that the detection of a product does not always mean a specific interaction has occurred between two regions. Therefore, a specific interaction between two regions is only confirmed when the interaction occurs at a higher frequency than with neighboring DNA.The ChIP-loop may be useful in identifying long-range cis-interactions and trans interaction mediated through proteins since frequent DNA collisions will not occur.
The 5C technique overcomes the junctional problems at the intramolecular ligation step and is useful for constructing complex interactions of specific loci of interest. This approach is unsuitable for conducting genome-wide complex interactions since that will require millions of 5C primers to be used.
In contrast to 3C and 5C, the 4C technique does not require the prior knowledge of both interacting chromosomal regions. Results obtained using 4C are highly reproducible with most of the interactions that are detected are between regions proximal to one another. On a single microarray, approximately a million interactions can be analyzed.
A common problem in all of these techniques is the requirement of a great number of cells, especially in the high-throughput methodologies. A single mammalian cell only provides two copies of any given restriction fragment, which can be ligated to only one other partner in 3C. Therefore any kind of quantitative analysis requires a large number of cells due to the need to determine the specification of an interaction between two regions. Experiments using the 4C technique routinely process ten million cells for analysis on a single microarray.
History
The 3C methodology was originally developed by Dekker and colleagues in 2002 at the University of MassachusettsUniversity of Massachusetts
This article relates to the statewide university system. For the flagship campus often referred to as "UMass", see University of Massachusetts Amherst...
and it aimed at identifying, locating and mapping physical interactions between genetic elements located throughout the human genome
Human genome
The human genome is the genome of Homo sapiens, which is stored on 23 chromosome pairs plus the small mitochondrial DNA. 22 of the 23 chromosomes are autosomal chromosome pairs, while the remaining pair is sex-determining...
. This technology would give beneficial insights into the complex interplay of genetic factors that contribute to such debilitating disorders such as cancer
Cancer
Cancer , known medically as a malignant neoplasm, is a large group of different diseases, all involving unregulated cell growth. In cancer, cells divide and grow uncontrollably, forming malignant tumors, and invade nearby parts of the body. The cancer may also spread to more distant parts of the...
, Duchenne muscular dystrophy
Duchenne muscular dystrophy
Duchenne muscular dystrophy is a recessive X-linked form of muscular dystrophy, which results in muscle degeneration, difficulty walking, breathing, and death. The incidence is 1 in 3,000 boys. Females and males are affected, though females are rarely affected and are more often carriers...
(DMD), Rett syndrome
Rett syndrome
Rett syndrome is a neurodevelopmental disorder of the grey matter of the brain that almost exclusively affects females. The clinical features include small hands and feet and a deceleration of the rate of head growth . Repetitive hand movements, such as wringing and/or repeatedly putting hands into...
and Alzheimer's disease
Alzheimer's disease
Alzheimer's disease also known in medical literature as Alzheimer disease is the most common form of dementia. There is no cure for the disease, which worsens as it progresses, and eventually leads to death...
.
3C is based on proximity ligation, which had been used previously to determine circularization frequencies of DNA in solution, and the effect of protein-mediated DNA bending on circularization. Seyfred and colleagues developed proximity ligation in nuclei: restriction enzyme digestion of unfixed nuclei and ligation in situ without diluting the chromatin, which they termed the "Nuclear Ligation Assay" (Cullen et al. Science. 1993 Jul 9;261(5118):203-6; Gothard LQ et al. Mol Endocrinol. 1996 Feb;10(2):185-95).

