DNA footprinting
Encyclopedia
DNA
footprinting is a method of investigating the sequence specificity of DNA-binding proteins in vitro. This technique can be used to study protein-DNA interactions both outside and within cells.
The regulation of transcription
has been studied extensively, and yet there is still much that is not known. Transcription factors and associated proteins that bind promoters, enhancers, or silencers
to drive or repress transcription are fundamental to understanding the unique regulation of individual genes
within the genome
. Techniques like DNA footprinting will help elucidate which proteins bind to these regions of DNA and unravel the complexities of transcriptional control.
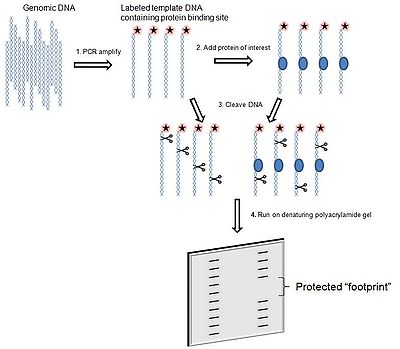 The simplest application of this technique is to assess whether a given protein binds to a region of interest within a DNA molecule. The wet lab methodology is summarized, with appropriate selection of reagents discussed, below.
The simplest application of this technique is to assess whether a given protein binds to a region of interest within a DNA molecule. The wet lab methodology is summarized, with appropriate selection of reagents discussed, below.
Note: Maxam-Gilbert chemical DNA sequencing
can be run alongside the samples on the polyacrylamide gel to allow the prediction of the exact location of ligand binding site.
DNA
Deoxyribonucleic acid is a nucleic acid that contains the genetic instructions used in the development and functioning of all known living organisms . The DNA segments that carry this genetic information are called genes, but other DNA sequences have structural purposes, or are involved in...
footprinting is a method of investigating the sequence specificity of DNA-binding proteins in vitro. This technique can be used to study protein-DNA interactions both outside and within cells.
The regulation of transcription
Transcription (genetics)
Transcription is the process of creating a complementary RNA copy of a sequence of DNA. Both RNA and DNA are nucleic acids, which use base pairs of nucleotides as a complementary language that can be converted back and forth from DNA to RNA by the action of the correct enzymes...
has been studied extensively, and yet there is still much that is not known. Transcription factors and associated proteins that bind promoters, enhancers, or silencers
Silencer (DNA)
In genetics a silencer is a DNA sequence capable of binding transcription regulation factors termed repressors. Upon binding, RNA polymerase is prevented from initiating transcription thus decreasing or fully suppressing RNA synthesis....
to drive or repress transcription are fundamental to understanding the unique regulation of individual genes
Gênes
Gênes is the name of a département of the First French Empire in present Italy, named after the city of Genoa. It was formed in 1805, when Napoleon Bonaparte occupied the Republic of Genoa. Its capital was Genoa, and it was divided in the arrondissements of Genoa, Bobbio, Novi Ligure, Tortona and...
within the genome
Genome
In modern molecular biology and genetics, the genome is the entirety of an organism's hereditary information. It is encoded either in DNA or, for many types of virus, in RNA. The genome includes both the genes and the non-coding sequences of the DNA/RNA....
. Techniques like DNA footprinting will help elucidate which proteins bind to these regions of DNA and unravel the complexities of transcriptional control.
Method

- Polymerase chain reactionPolymerase chain reactionThe polymerase chain reaction is a scientific technique in molecular biology to amplify a single or a few copies of a piece of DNA across several orders of magnitude, generating thousands to millions of copies of a particular DNA sequence....
(PCR) amplify and label region of interest that contains a potential protein-binding site, ideally amplicon is between 50 to 200 base pairs in length. - Add protein of interest to a portion of the labeled template DNA; a portion should remain separate without protein, for later comparison
- Add a cleavage agent to both portions of DNA template. The cleavage agent is a chemical or enzyme that will cut at random locations in a sequence independent manner. The reaction should occur just long enough to cut each DNA molecule in only one location. A protein that specifically binds a region within the DNA template will protect the DNA it is bound to from the cleavage agent.
- Run both samples side by side on a polyacrylamidePolyacrylamidePolyacrylamide is a polymer formed from acrylamide subunits. It can be synthesized as a simple linear-chain structure or cross-linked, typically using N,N-methylenebisacrylamide. Polyacrylamide is not toxic...
gel electrophoresisGel electrophoresisGel electrophoresis is a method used in clinical chemistry to separate proteins by charge and or size and in biochemistry and molecular biology to separate a mixed population of DNA and RNA fragments by length, to estimate the size of DNA and RNA fragments or to separate proteins by charge...
. The portion of DNA template without protein will be cut at random locations, and thus when it is run on a gel, will produce a ladder-like distribution. The DNA template with the protein will result in ladder distribution with a break in it, the "footprint", where the DNA has been protected from the cleavage agent.
Note: Maxam-Gilbert chemical DNA sequencing
DNA sequencing
DNA sequencing includes several methods and technologies that are used for determining the order of the nucleotide bases—adenine, guanine, cytosine, and thymine—in a molecule of DNA....
can be run alongside the samples on the polyacrylamide gel to allow the prediction of the exact location of ligand binding site.
Labeling
The DNA template can be labeled at the 3' or 5' end, depending on the location of the binding site(s). Labels that can be used are:- Radioactivity has been traditionally used to label DNA fragments for footprinting analysis, as the method was originally developed from the Maxam-Gilbert chemical sequencing technique. Radioactive labeling is very sensitive and is optimal for visualizing small amounts of DNA.
- FluorescenceFluorescenceFluorescence is the emission of light by a substance that has absorbed light or other electromagnetic radiation of a different wavelength. It is a form of luminescence. In most cases, emitted light has a longer wavelength, and therefore lower energy, than the absorbed radiation...
is a desirable advancement due to the hazards of using radio-chemicals. However, it has been more difficult to optimize because it is not always sensitive enough to detect the low concentrations of the target DNA strands used in DNA footprinting experiments. Electrophoretic sequencing gels or capillary electrophoresisCapillary electrophoresisCapillary electrophoresis , also known as capillary zone electrophoresis , can be used to separate ionic species by their charge and frictional forces and hydrodynamic radius. In traditional electrophoresis, electrically charged analytes move in a conductive liquid medium under the influence of an...
have been successful in analyzing footprinting of fluorescently tagged fragments.
Cleavage agent
A variety of cleavage agents can be chosen. Ideally a desirable agent is one that is sequence neutral, easy to use, and is easy to control. Unfortunately none available meet all these all of these standards, so an appropriate agent can be chosen, depending on your DNA sequence and ligand of interest. The following cleavage agents are described in detail:- DNase I: a large protein that functions as a double-strand endonucleaseEndonucleaseEndonucleases are enzymes that cleave the phosphodiester bond within a polynucleotide chain, in contrast to exonucleases, which cleave phosphodiester bonds at the end of a polynucleotide chain. Typically, a restriction site will be a palindromic sequence four to six nucleotides long. Most...
. It binds the minor groove of DNA and cleaves the phosphodiester backbone. It is a good cleavage agent for footprinting because its size makes it easily physically hindered. Thus is more likely to have its action blocked by a bound protein on a DNA sequence. In addition, the DNase I enzyme is easily controlled by adding EDTA to stop the reaction. There are however some limitations in using DNase I. The enzyme does not cut DNA randomly; its activity is affected by local DNA structure and sequence and therefore results in an uneven ladder. This can limit the precision of predicting a protein’s binding site on the DNA molecule. - Hydroxyl radicals: are created from the Fenton reaction, which involves reducing Fe2+ with H2O2 to form free hydroxyl molecules. These hydroxyl molecules react with the DNA backbone, resulting in a break. Due to their small size, the resulting DNA footprint has high resolution. Unlike DNase I they have no sequence dependence and result in a much more evenly distributed ladder. The negative aspect of using hydroxyl radicals is that they are more time consuming to use, due to a slower reaction and digestion time.
- Ultraviolet irradiation: can be used to excite nucleic acids and create photoreactions, which results in damaged bases in the DNA strand. Photoreactions can include: single strand breaks, interactions between or within DNA strands, reactions with solvents, or crosslinks with proteins.
- The workflow for this method has an additional step, once both your protected and unprotected DNA have been treated, there is subsequent primer extension of the cleaved products. The extension will terminate upon reaching a damaged base, and thus when the PCR products are run side-by-side on a gel; the protected sample will show an additional band where the DNA was crosslinked with a bound protein.
- Advantages of using UV are that it reacts very quickly and can therefore capture interactions that are only momentary. Additionally it can be applied to in vivoIn vivoIn vivo is experimentation using a whole, living organism as opposed to a partial or dead organism, or an in vitro controlled environment. Animal testing and clinical trials are two forms of in vivo research...
experiments, because UV can penetrate cell membranes. A disadvantage is that the gel can be difficult to interpret, as the bound protein does not protect the DNA, it merely alters the photoreactions in the vicinity.
In vivo footprinting
- In vivo footprinting is a technique used to analyze the protein-DNA interactions that are occurring in a cell at a given time point. DNase I can be used as a cleavage agent if the cellular membrane has been permeabilized. However the most common cleavage agent used is UV irradiation because it penetrates the cell membrane without disrupting cell state and can thus capture interactions that are sensitive to cellular changes. Once the DNA has been cleaved or damaged by UV, the cells can be lysed and DNA purified for analysis of a region of interest.
- Ligation-mediated PCR is an alternative method to footprint in vivo. Once a cleavage agent has been used on the genomic DNA, resulting in single strand breaks, and the DNA is isolated, a linker is added on to the break points. A region of interest is amplified between the linker and a gene-specific primer, and when run on a polyacrylamide gel, will have a footprint where a protein was bound.
- In vivo footprinting combined with immunoprecipitationImmunoprecipitationImmunoprecipitation is the technique of precipitating a protein antigen out of solution using an antibody that specifically binds to that particular protein. This process can be used to isolate and concentrate a particular protein from a sample containing many thousands of different proteins...
can be used to assess protein specificity at many locations throughout the genome. The DNA bound to a protein of interest can be immunoprecipitated with an antibody to that protein, and then specific region binding can be assessed using the DNA footprinting technique.
Quantitative footprinting
- The DNA footprinting technique can be modified to assess the binding strength of a protein to a region of DNA. Using varying concentrations of the protein for the footprinting experiment, the appearance of the footprint can be observed as the concentrations increase and the proteins binding affinity can then be estimated.

