Polymerase cycling assembly
Encyclopedia
Polymerase cycling assembly (or PCA, also known as Assembly PCR) is a method for the assembly of large DNA
oligonucleotide
s from shorter fragments. The process uses the same technology as PCR
, but takes advantage of DNA hybridization and annealing as well as DNA polymerase
to amplify a complete sequence of DNA in a precise order based on the single stranded oligonucleotides used in the process. It thus allows for the production of synthetic genes and even entire synthetic genome
s.
 Much like how primers are designed such that there is a forward primer and a reverse primer capable of allowing DNA polymerase to fill the entire template sequence, PCA uses the same technology but with multiple oligonucleotides. While in PCR the customary size of oligonuleotides used is 18 base pairs, in PCA lengths of up to 50 are used to ensure uniqueness and correct hybridization.
Much like how primers are designed such that there is a forward primer and a reverse primer capable of allowing DNA polymerase to fill the entire template sequence, PCA uses the same technology but with multiple oligonucleotides. While in PCR the customary size of oligonuleotides used is 18 base pairs, in PCA lengths of up to 50 are used to ensure uniqueness and correct hybridization.
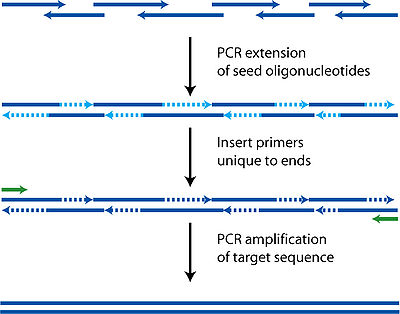
Each oligonucleotide is designed to be either part of the top or bottom strand of the target sequence. As well as the basic requirement of having to be able to tile the entire target sequence, these oligonucleotides must also have the usual properties of similar melting temperatures, hairpin free, and not too GC rich to avoid the same complications as PCR.
During the polymerase cycles, the oligonucleotides anneal to complementary fragments and then are filled in by polymerase. Each cycle thus increases the length of various fragments randomly depending on which oligonucleotides find each other. It is critical that there is complementarity between all the fragments in some way or a final complete sequence will not be produced as polymerase requires a template to follow.
After this initial construction phase, additional primers encompassing both ends are added to perform a regular PCR reaction, amplifying the target sequence away from all the shorter incomplete fragments. A gel purification can then be used to identify and isolate the complete sequence.
A modification of this method, Gibson assembly
, described by Gibson et al. allows for single-step isothermal assembly of DNA with up to several hundreds kb. By using T5 exonuclease to 'chew back' complementary ends, an overlap of about 40bp can be created. The reaction takes place at 50°C, a temperature where the T5 exonuclease is unstable. After a short timestep it is degraded, the overlapps can anneal and be ligated. Cambridge University IGEM
team made a video describing the process.
DNA
Deoxyribonucleic acid is a nucleic acid that contains the genetic instructions used in the development and functioning of all known living organisms . The DNA segments that carry this genetic information are called genes, but other DNA sequences have structural purposes, or are involved in...
oligonucleotide
Oligonucleotide
An oligonucleotide is a short nucleic acid polymer, typically with fifty or fewer bases. Although they can be formed by bond cleavage of longer segments, they are now more commonly synthesized, in a sequence-specific manner, from individual nucleoside phosphoramidites...
s from shorter fragments. The process uses the same technology as PCR
Polymerase chain reaction
The polymerase chain reaction is a scientific technique in molecular biology to amplify a single or a few copies of a piece of DNA across several orders of magnitude, generating thousands to millions of copies of a particular DNA sequence....
, but takes advantage of DNA hybridization and annealing as well as DNA polymerase
DNA polymerase
A DNA polymerase is an enzyme that helps catalyze in the polymerization of deoxyribonucleotides into a DNA strand. DNA polymerases are best known for their feedback role in DNA replication, in which the polymerase "reads" an intact DNA strand as a template and uses it to synthesize the new strand....
to amplify a complete sequence of DNA in a precise order based on the single stranded oligonucleotides used in the process. It thus allows for the production of synthetic genes and even entire synthetic genome
Genome
In modern molecular biology and genetics, the genome is the entirety of an organism's hereditary information. It is encoded either in DNA or, for many types of virus, in RNA. The genome includes both the genes and the non-coding sequences of the DNA/RNA....
s.
PCA principles

Each oligonucleotide is designed to be either part of the top or bottom strand of the target sequence. As well as the basic requirement of having to be able to tile the entire target sequence, these oligonucleotides must also have the usual properties of similar melting temperatures, hairpin free, and not too GC rich to avoid the same complications as PCR.
During the polymerase cycles, the oligonucleotides anneal to complementary fragments and then are filled in by polymerase. Each cycle thus increases the length of various fragments randomly depending on which oligonucleotides find each other. It is critical that there is complementarity between all the fragments in some way or a final complete sequence will not be produced as polymerase requires a template to follow.
After this initial construction phase, additional primers encompassing both ends are added to perform a regular PCR reaction, amplifying the target sequence away from all the shorter incomplete fragments. A gel purification can then be used to identify and isolate the complete sequence.
A modification of this method, Gibson assembly
Gibson assembly
Gibson assembly is a DNA assembly method which allows for the joining of multiple DNA fragments in a single, isothermal reaction. It was invented in 2009 by the Daniel Gibson while he was at the J...
, described by Gibson et al. allows for single-step isothermal assembly of DNA with up to several hundreds kb. By using T5 exonuclease to 'chew back' complementary ends, an overlap of about 40bp can be created. The reaction takes place at 50°C, a temperature where the T5 exonuclease is unstable. After a short timestep it is degraded, the overlapps can anneal and be ligated. Cambridge University IGEM
IGEM
The International Genetically Engineered Machine competition is a worldwide Synthetic Biology competition aimed at undergraduate university students.- Competition details :...
team made a video describing the process.

